Today’s guest blog is written by Kiley Roach, Zymo Research
Autosomal short tandem repeat (STR) analysis has become the gold standard for profiling in the field of forensic science. However, due to the limitations of sample quantity, polymerase chain reaction (PCR) technology is used to amplify samples prior to STR analysis. As a result, STR profiling can become highly problematic if the initial sample contains compounds that inhibit PCR. These polyphenolic PCR inhibitors are organic compounds found in the environment and bodily fluids and are often problematic even at very low concentrations. Some of these inhibitors block the DNA polymerase binding site and inhibit amplification, whereas others directly interact with the DNA polymerase or it’s cofactor, thus reducing PCR amplification efficiency. The reduced amplification efficiency interferes with the multiplex STR profiling, especially with samples containing low concentrations of DNA. Thus, a highly effective PCR inhibitor removal system that can be easily integrated into current DNA extraction workflows is crucial to ensure PCR amplification is successful and STR analysis is accurate.
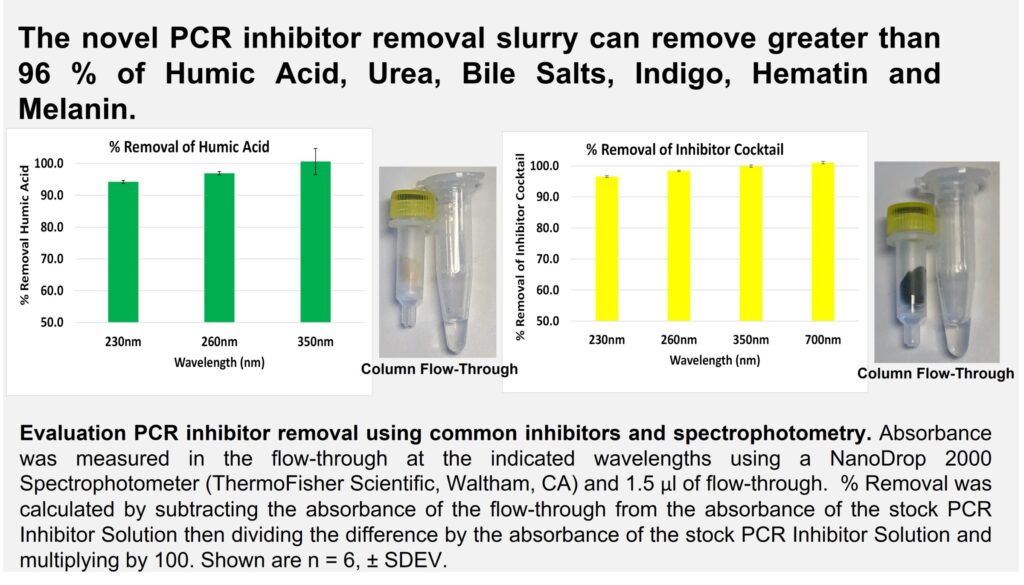
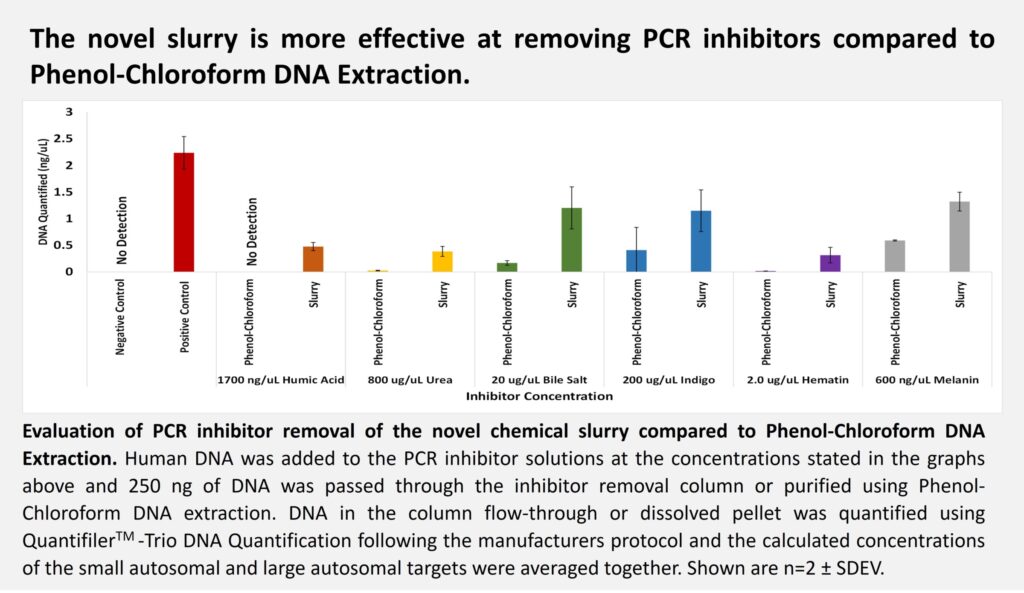
We presented a novel chemical slurry at ISHI 2022 that we developed for removing PCR inhibitors after nucleic acid extraction. The effectiveness of our PCR inhibitor removal slurry was evaluated by passing human DNA spiked with either humic acid, urea, bile salts, indigo, hematin or melanin through a spin-column containing the slurry. PCR inhibitor removal was then assessed in the flow through by measuring inhibitor concentration using spectrophotometry and PCR inhibition using the QuantifilerTM Trio DNA Quantification Kit. Additionally, DNA spiked with inhibitors was also processed using a traditional phenol-chloroform extraction protocol in parallel for comparison.
The absorption spectrum data revealed that the slurry was able to remove greater than 96% of all the inhibitors tested (see figure 1). Furthermore, we also successfully demonstrated that the slurry could restore PCR amplification of contaminated samples and is more effective at removing PCR inhibitors than a traditional phenol-chloroform extraction protocol (see figure 2). Based on these results, we established that our novel chemical slurry is an easy and efficient method for removing PCR inhibitors.
Our team became interested in this work after consulting with multiple forensics labs and learning that PCR inhibitors prove to be a relevant problem in the field. Many small-scale or second opinion labs work with complex samples that contain contaminants from the surrounding environment which require diluting and/or multiple rounds of phenol-chloroform extractions before the QuantifilerTM Trio system can quantify the DNA. This, as you can imagine, is extremely time consuming and can be problematic if the samples have low DNA yield to begin with. Our goal was to come up with an easy solution that would save time and work for low DNA input samples.
The simplicity of our PCR inhibitor removal slurry revolutionizes sample clean-up for forensics analysis. After just 1 minute of centrifugation, samples are free from any inhibitors that could be detrimental to PCR amplification. This will help forensics labs save both time and resources. Instead of using specially trained lab technicians for phenol-chloroform extractions, which take a considerable amount of time, reagents, and equipment, labs can use our inhibitor removal slurry to consistently achieve cleaner samples with a simple centrifugation step. Additionally, the inhibitor removal slurry can be used after any purification kit. This means that forensics labs would not need to change any of their existing DNA extraction methods in order to implement this product.
Our goal from attending ISHI was to make attendees aware of this novel PCR inhibitor removal slurry and further gauge the need in the field for such a tool. After speaking with multiple attendees about their work, it became apparent that there is a clear split in the field regarding PCR inhibitors. For companies working with complex samples collected in the field, such as unidentified remains, PCR inhibitors are a significant problem, whereas companies working with samples collected using a highly controlled method, such as buccal swabs, PCR inhibitors are not as problematic. From speaking with people from both sides of the spectrum, we learned that for this tool to be adopted by the forensics field, we would need to ensure that the product is free from human DNA.
Moving forward, we are going to perform STR analysis on the samples that were processed with our PCR inhibitor removal slurry. We have already proven that our chemical slurry is successful in restoring DNA quantification using the QuantifilerTM Trio kit, but we still need to see how our PCR inhibitor removal system impacts the resulting STR profile. Additionally, we already offer a complete line of DNA purification products for a variety of sample types, therefore, we are also planning on evaluating our DNA purification kits for STR analysis. With this, we could offer an all-inclusive DNA extraction workflow that guarantees forensics samples are free from any PCR inhibitors.
If you are interested in testing out our PCR inhibitor removal slurry in your current DNA profiling workflow, we do have free samples available. Please email kroach@zymoresearch.com with your shipping address to receive your free sample.
WOULD YOU LIKE TO SEE MORE ARTICLES LIKE THIS? SUBSCRIBE TO THE ISHI BLOG BELOW!
SUBSCRIBE NOW!