Signature Science’s Center for Advanced Genomics® (CAG) is working to identify and assess the applicability of emerging technologies and methodologies for use in forensic DNA analyses. To this end, the CAG has validated array-based SNP genotyping using the lllumina® Global Screening Array (GSA) for use on forensic samples. As the demand for forensic genetic genealogy (FGG) services has emerged since the 2018 identification of Joseph DeAngelo as the Golden State Killer, most FGG providers have outsourced cold case samples to clinical laboratories for SNP array methods but, in these testing environments, high quality and quantities of DNA are optimal. The goal of our developmental validation was to study and characterize the applicability of array-based SNP genotyping to challenging forensic sample types.
Written by: David Russell, Signature Science
Validation Study and Results. The CAG’s developmental validation was guided by FBI QAS and SWGDAM standards with the intended application to challenging forensic samples. The studies performed [sensitivity, precision and accuracy (repeatability and reproducibility), mixtures, degradation, species specificity, contamination, and case-type samples] utilized extensively characterized human genomes for which high-confidence variant calls are known. Concordance was assessed using the NIST/Genome-in-a-bottle (GIAB) sample call sets as truth data.
The Infinium® assay workflow is a genome-wide microarray genotyping assay that utilizes the BeadChip platform.1 This accurate and flexible microarray technology allows for the interrogation of a large number of SNPs through unlimited loci multiplexing.2–4 Through our validation efforts, the CAG was able to demonstrate sensitivity at 0.2 ng of DNA input (significantly reduced from the manufacturer’s recommended 200 ng input), which is consistent with most forensic samples. The precision and accuracy (repeatability and reproducibility) studies showed average concordance rates at 99.2% across all samples.
Repeatability and reproducibility studies demonstrated reliable and consistent call rates and high concordance, regardless of operator.5 These key studies give confidence in the method’s ability to accurately genotype samples with low DNA inputs. In the mixture study profile generated from samples at a mixture ratio of 3:1 or greater between the major and minor contributors, the resulting major contributor’s profile is accurate, showing 98.85% (3:1) – 99.99% (9:1) concordance.6 This information demonstrated that under certain mixture proportions the genotype of the unknown major contributor can be determined, with high accuracy, without removing the known contributor. Mixture deconvolution with array data is considerably difficult and currently not standard practice for our services. However, with this knowledge, it will allow us to assess the data further and make recommendations on how to proceed with processing the case.
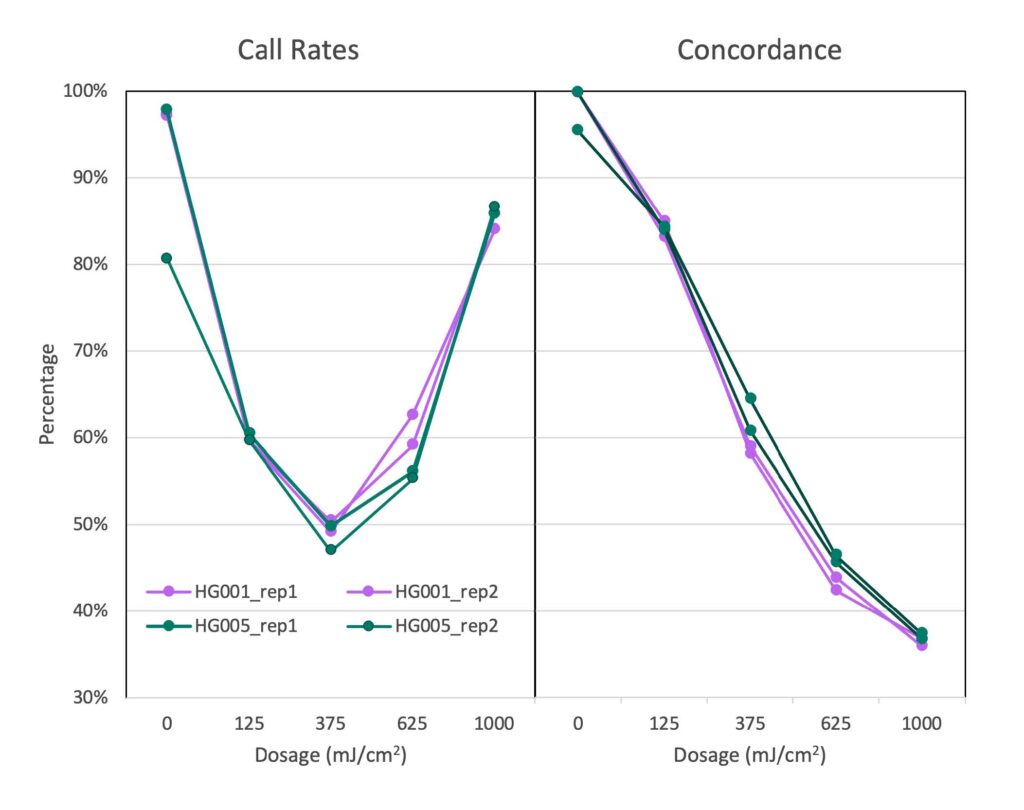
The data generated during the degradation study was unexpected; samples known to have been severely degraded produced genotyping data with call rates similar to pristine samples.7 Lower heterozygosity was observed in the degraded samples (average of 7%) compared to the GIAB population set (average of 17.3%). This suggests that allelic dropout is responsible for the noticeable shift to false homozygous calls, accounting for the high call rate yet discordant SNP data. Overall, we observed that call rate alone was not a reliable indicator of genotyping success; therefore, more metrics were investigated to assess sample quality and guide sample processing.
Conclusion. This validation characterized the performance of forensic samples on the GSA platform, providing valuable insight into what forensic practitioners can expect from microarray-based genotyping data. Thresholds for call rate, heterozygosity, and overall signal (fluorescence) were established to assess the data and determine suitability of samples for the workflow. Ultimately, this work should provide a better understanding of how microarray systems will operate in the forensic community as the application of this technology to more cases continues to grow.
What’s Next? The CAG is currently evaluating next generation sequencing methods to support FGG services and is validating the Verogen ForenSeq® Kintelligence workflow for the NDIS-approved MiSeq FGx® sequencing system.
More information. Posters presented on this validation are available through FigShare or in the references listed. For more information about Signature Science, the Center for Advanced Genomics, and our FGG services visit our website or contact David Russell.
About the Author David Russell, M.S. is a Unit Manager and Forensic Scientist for Signature Science’s Center for Advanced Genomics. He leads research, development, testing, and evaluation efforts. His technical forensic expertise includes forensic DNA STR analysis and interpretation, MPS library preparation, and array-based SNP genotyping workflows. Mr. Russell earned his M.S. in Biology (Microbial Forensics) from Western Carolina University.
References:
- Gunderson KL, Steemers FJ, Ren H, Ng P, Zhou L, Tsan C, Chang W, Bullis D, Musmacker J, King C, et al. Whole‐Genome Genotyping. In: Methods in Enzymology. Vol. 410. Elsevier; 2006. p. 359–376. https://linkinghub.elsevier.com/retrieve/pii/S0076687906100178. doi:10.1016/S0076-6879(06)10017-8
- Fan J, Gunderson KL, Bibikova M, Yeakley JM, Chen J, Wickham Garcia E, Lebruska LL, Laurent M, Shen R, Barker D. [3] Illumina Universal Bead Arrays. In: Methods in Enzymology. Vol. 410. Elsevier; 2006. p. 57–73. https://linkinghub.elsevier.com/retrieve/pii/S0076687906100038. doi:10.1016/S0076-6879(06)10003-8
- Steemers FJ, Chang W, Lee G, Barker DL, Shen R, Gunderson KL. Whole-genome genotyping with the single-base extension assay. Nature Methods. 2006;3(1):31–33. doi:10.1038/nmeth842
- Illumina. Infinium Assay Workflow. 2012. (Technology Spotlight: SNP genotyping).
- Russell D, Elayna Moreithi, Neal C, Heaton M, Turner S, Reedy C. Optimizing Sensitivity and Validating the Illumina Infinium Assay for Genotyping of Forensically Relevant Sample Types for Investigative Lead Generation. figshare; 2020. p. 1167388 Bytes. https://figshare.com/articles/Optimizing_Sensitivity_and_Validating_the_Illumina_Infinium_Assay_for_Genotyping_of_Forensically_Relevant_Sample_Types_for_Investigative_Lead_Generation/11808939/2. doi:10.6084/M9.FIGSHARE.11808939.V2
- Russell, David, Reedy, Carmen, Turner, Stephen, Guertin, Stephanie, Heaton, Mary, Bouchet, Jessica, Peck, Michelle, Gorden, Erin, Ciuzio, Elayna, Neal, Christina. Developmental Validation of the Illumina Infinium Assay using the Global Screening Array (GSA) on the iScan System for use in Forensic Laboratories. figshare; 2021. p. 313480 Bytes. https://figshare.com/articles/poster/Developmental_Validation_of_the_Illumina_Infinium_Assay_using_the_Global_Screening_Array_GSA_on_the_iScan_System_for_use_in_Forensic_Laboratories/16589771/2. doi:10.6084/M9.FIGSHARE.16589771.V2
- Russell D, Reedy C, Elayna Moreithi, Neal C, Turner S, Heaton M. Evaluation of Degraded Human DNA Samples Using the Illumina® Global Screening Array. figshare; 2021. p. 1084147 Bytes. https://figshare.com/articles/poster/Evaluation_of_Degraded_Human_DNA_Samples_Using_the_Illumina_Global_Screening_Array/13582181/3. doi:10.6084/M9.FIGSHARE.13582181.V3
WOULD YOU LIKE TO SEE MORE ARTICLES LIKE THIS? SUBSCRIBE TO THE ISHI BLOG BELOW!
SUBSCRIBE NOW!


