Today’s guest blog is written by Michael Marciano, Ph.D and Amber Vandepoele, Syracuse University
Mixture interpretation remains a central challenge in the forensic DNA field. Much research and development has focused on methods to improve the interpretation of complex samples including software solutions. These methods are used to improve mixture interpretation on the “back-end” rather than address the separation of the individual biological components (e.g., cells) to wholly avoid mixtures. This project will address this need in the forensic biology field, aiming to develop and optimize a method to identify and recover male cells from a mixture of male and female like-cells, e.g., epithelial cells. For example, the identification of a vasectomized male in a sexual assault of a female victim. The method will adapt two well characterized methods into a single unified protocol for use in forensic DNA analyses – Y chromosome targeting via the Abbott Molecular Vysis CEP Y DYZ1 probe and male cell recovery using the DEPArray NxT or PLUS. Separating male from female cells in like-cell mixtures will result in single source male autosomal profiles and provide unprecedented levels of resolution that will lead to stronger statistical support for the comparisons. This study is in progress.
The lab group evaluated the DEPArray in a previous NIJ project (2015-NE-BX-K002) and the manufacturer, (Menarini) put us in touch with Jeremy Dubois at the Acadiana Criminalistics Laboratory in Louisiana. Jeremy believed that this technology could be used to address casework that involves vasectomized male perpetrators of sexual assaults. We partnered with Jeremy, director Kevin Ardoin, and the Acadiana lab to propose this study which was later awarded funding by the NIJ to investigate this methodology (Award: 2020-DQ-BX-0019).
The proposed project will address this critical need in the forensic biology field, aiming to evaluate an emerging method that will significantly improve results and conclusions for a specific case type, improve laboratory efficiency, and lower the cost of processing such cases. Separating male from female cells in like-cell mixtures will result in single source male autosomal profiles and provide unprecedented levels of resolution that will lead to stronger statistical support for the analysts’ results. It may also allow the profiles to be uploaded to CODIS in cases that yielded profiles that were not previously able to be uploaded. The proposed method would also simplify the data interpretation, as probabilistic genotyping would not have to be used to deconvolute a mixture. The benefits extend to any sample with a male and female mixture of like-cells. For example, this method could be used in a case where a knife was collected from a female homicide victim’s home. The majority of cells on the knife would be contributed by the female victim, however this method could separate male from female cells improving the chances of success.
First, a Y-Chromosome staining procedure was developed. The Abbott CEP Y Spectrum Green and Orange probes were used to selectively label the Yq12 satellite III region of the Human Y chromosome. The manufacturer’s recommended protocol was adapted and optimized for this application (Figure 2).

Next, staining efficiency was evaluated by testing the stain on cultured cells (male-RWPE prostate cells and female-HeLa cells) on a slide. This was followed by evaluating the staining method in solution (a requirement for DEPArray mediated detection and recovery. The optimization of the method in solution included testing the efficacy of detergents (Triton, NP-40), internally made denaturation solution and Abbott hybridization buffer, 3:1 methanol/acetic acid and 2% PFA, incubation times – 1 hr., 3 hr. and overnight.
After staining, the Y-probe stained male cells are identified recovered using the DEPArray NxT and PLUS (Figure 4).
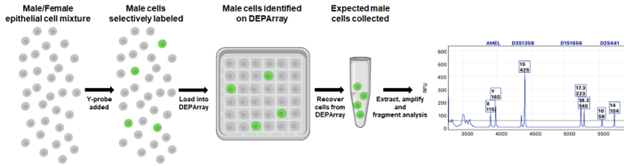
Manufacturer recommended protocols were used for sample preparation and instrument protocols. Cells were routed and recovered in two primary groups (1) single cells – male or female and (2) groups of cells – Y-probe positive cells or Y-probe positive cells and Y-probe negative (expected male and female cells in a single recovery). Samples were extracted using the Menarini LYSE Prep kit (DALYS) and amplified using PowerPlex Fusion 6c Human DNA amplification kit (Promega Corporation) and fragments were detected using the 3500xL Genetic Analyzer (ThermoFisher Scientific) – all procedures were performed using the manufacturer’s recommended parameters.
Staining of the cultured RWPE (male) and HeLa (female) cells demonstrated the effectiveness of the staining method. Green Y-probe signal was seen in RWPE cells but not Hela cells. Staining of the male and female buccal epithelial cells with the green probe resulted in Y-probe signal in male cells and no Y-probe signal in female cells with staining efficiencies of 68.76% ± 28.66 and 0.48% ± 0.28%, respectively. Next, the DEPArray NxT was used to separate a mixture of male and female buccal cells stained with the green probe. The green probe was difficult to visualize in the FITC channel, therefore, the orange probe (PE channel) was evaluated and found to provide better quality signal, with a staining efficiency of 75.41% ± 13.88% and 0.48% ± 0.28% for male and female buccal epithelial cells, respectively.
Samples stained with the Y-probe were loaded onto the DEPArray NxT or PLUS. The ability to identify and recover male (Y-probe stained) and female cells (unstained) on the DEPArray was first assessed through the recovery of single cells followed by profiling. 80% of expected male single cells generated a male profile (true positive) and 79% of expected female single cells generated a female profile (true negative). The false positives may be a result of high background, a threshold is being investigated to avoid this. The false negative results were consistent with what was observed in the staining efficiency study. A total of 7 mixtures runs were carried out with male to female dilutions of 1:1 (2) and 1:10 (5). Genetic profiles of expected male cells recovered from the DEPArray were evaluated based on mean peak heights and the proportion of alleles present for the male and female donors. Single source male profiles were obtained from both 1:1 mixtures as well as three 1:10 mixtures. Two 1:10 mixtures resulted in mixed profiles with ratios of 1.1:1 and 0.85:1 male to female cells, respectively. Additional work is underway including the analysis of samples with higher ratios of female to male cells.
This in progress study has demonstrated a method to stain human Y-chromosomes in solution and recover male cells with a success rate of 80%. In addition, the collection of male cells when recovered in groups has shown to be successful in 1:1 and 1:10 dilutions of male to female epithelial cells. In only a single case were female alleles detected. Despite this, an improvement in the male to female ratio was observed and, therefore, still increased the ability to successfully deconvolute the mixture. Additional work is ongoing to further test dilutions of up to 1 to 100 (male to female) and continue characterizing the method on mixtures of lower quality, “old” cells.
Many key collaborators allowed for the success of this project. The research was primarily conducted by Jonathan Hogg and Amber C. W. Vandepoele with support from Nori Zaccheo, Morgan Frank, and Haley Crooks at the Syracuse University Forensic and National Security Sciences Institute. Additional guidance and support on the project were by Janine Schulte and Dr. iris Schulz from the Institute of Forensic Medicine at the University of Basel and Jeremy Dubois and Kevin Ardoin from the Acadiana Criminalistics Laboratory.
WOULD YOU LIKE TO SEE MORE ARTICLES LIKE THIS? SUBSCRIBE TO THE ISHI BLOG BELOW!
SUBSCRIBE NOW!


