Today’s blog is written by guest bloggers, Lutz Roewer and Sascha Willuweit, Institute of Legal Medicine and Forensic Sciences, Dept. Forensic Genetics, Berlin, Germany. Reposted from The ISHI Report with permission.
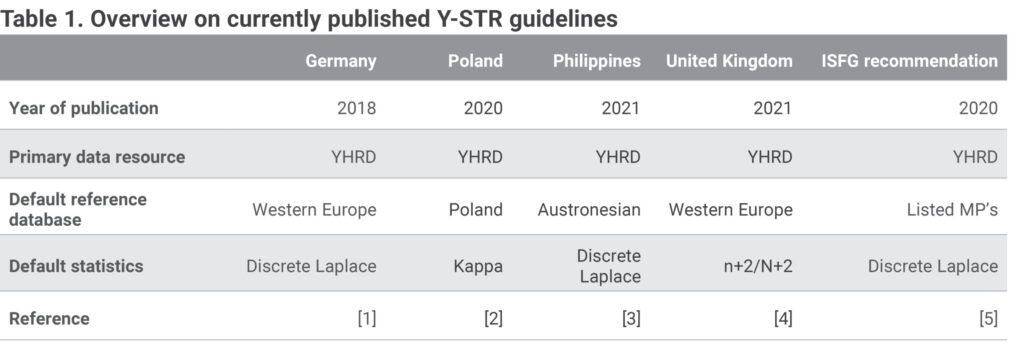
The reporting of Y-STR results with meaningful statistics requires a look beyond the borders, borders of the standard forensic mathematics practiced for decades and political borders. To substantiate this view, let us consult some recently published regulations on forensic Y chromosome testing and statistical interpretation of Y-STR results, for example of Germany [1], Poland [2], the Philippines [3] and the United Kingdom [4] (Table 1). In the United States, the Y-STR interpretation guidelines traditionally provided by the Scientific Working Group on DNA Analysis Methods (SWGDAM) are nearly finalized. Other countries follow the ISFG recommendations published recently [5]. A closer look at these documents reveals the different use of reference data and statistical methods. In this brief correspondence, we want to touch on the challenging problem of how global data resources and statistical methods based on an evolutionary genetic model, which are specifically designed for Y-STR interpretation can be incorporated in a workable approach that forensic experts in a country can follow in their daily routine.
The expert panels on Y-STR testing convened in different countries agree to recommend the global YHRD database (https://yhrd.org) as the primary source of reference data, however they disagree on the best statistical model to calculate haplotype frequencies. The result is a heterogeneous picture of methods. While the Discrete Laplace (DL) calculation [6], endorsed by the ISFG [5], is recommended as the default method for example in Germany and the Philippines, other countries consider different modifications of the counting method more appropriate. Poland recommends the Kappa method [7] and the United Kingdom the augmented count estimate n+2/N+2 [8]. In addition, the recommendations on the choice of the default reference databases within the YHRD differ remarkably. Some countries (Germany, United Kingdom, Philippines) prefer metapopulations i.e. groups of population samples shown to have characteristic common features (e.g. language, demography, phylogenetic SNPs) while others prefer national databases (in Poland).
No doubt, all these methods and databases have limitations. The limitation of different counting methods is the direct relationship between the evidentiary weight and the size of the database [2]. On the other hand, the count estimates are easy to defend at court and are seen as “conservative”. This is not the case for the extrapolation (Discrete Laplace, DL) method, which requires a number of assumptions on the composition of the reference population(s) and the evolution of haplotypes by mutation. The method is much more independent from the database size and “realistic”. The DL method has its limitations in the calculation of partial profiles or profiles bearing non-consensus alleles. The calculation for larger STR panels requires considerable computer power. In addition, calculations are prone to errors if a database consists of subpopulations. Currently the DL method is enabled at YHRD and provides metapopulation-based calculations for core haplotypes (17 loci) included in widely used commercial kits.
The DL method using the full global YHRD dataset extracts important information from a haplotype match, which must be given special consideration by the forensic expert. Let’s demonstrate this with an example:
In a forensic investigation a Y-STR haplotype is found identical in a trace and a suspect. The haplotype is observed Zero times in a national Y chromosome reference database (N = 15,000). This database is composed of five defined subpopulations each of which has N = 3000. The observed frequency using the augmented counting method (n+1/N+1) would be 1/15,001 in the general database and 1/3,001 in all five subpopulation databases. If guidelines stipulate the reporting of count results, the qualitative statement “non-exclusion of the suspect as the donor of the trace” would be further specified using these five frequency values, which describe the likelihood that the matching haplotype occurs by chance in any of the subpopulations. The higher the frequency of the haplotype in the reference database, the more it relieves the defendant. In our example, the value of the Y-STR evidence is extremely low. And this is not because of the incapability of the method, but due to the small numbers of reference samples on the one hand and the inadequacy of the estimation method on the other. Forensic genetic research recently succeeded to show, that another grouping principle for reference populations and a statistical method that can analyze haplotype distributions, could significantly increase the evidential value of a trace/person match based exclusively on Y-STRs.
The YHRD is structured in a way, that new categories of metapopulation (MP) databases (larger than national) are built on basis of ancestry groups defined by geography, demography, language and phylogenetic Y-SNP information [9]. These MP’s include population samples that are related and distributed globally. Names for these new categories have been found which differ from national census schemes that often follow historical conventions. A good example for such a historical name is “Caucasian”. In the YHRD samples categorized as US Caucasians belong to the European MP. The YHRD further provides a number of statistical methods built upon haplotype distributions in populations, among these the Discrete Laplace method [9]. This method is able to analyze the connectivity of haplotypes in a metapopulation if it is built on basis of common ancestry and represents the prevailing deep-rooting male lineages. The probability of occurrence of a haplotype, whether it is observed or not in this and any other MP, can then be estimated.
Now, if the frequency of the haplotype of our example is calculated using DL in different MP’s, there will be a number of highly diverging estimates. This means, by chance the haplotype was not sampled in the five rather small national databases, but is nonetheless rather frequent in one or more worldwide populations and rare in others. Among these populations is probably the one, which represents the ancestors of the trace donor.
Regardless of an intriguing result like this, the true donor of the trace remains unknown, because the Y-STR method does not disclose the individuality of the DNA. However, a naïve counting in the five example databases resulting in equal values dismissed the information, which is built-in in the haplotype structure and accessible via the DL method and global databases. The judgement of the count and the DL outcome is the exclusive responsibility of the court, but the available information (if requested) should not be withheld. The availability and scrutiny of representative reference data placing the evidence in a context is a strong argument for the applicability of the Y-STR method in casework.
To summarize: We think that with sufficient empirical data, well structured databases and a suitable statistics the court-going forensic experts can prove their expertise on Y-STR interpretation, with defensive methods like counting (naive, augmented, kappa inflated) their expertise for using Y-STRs can be severely contested.
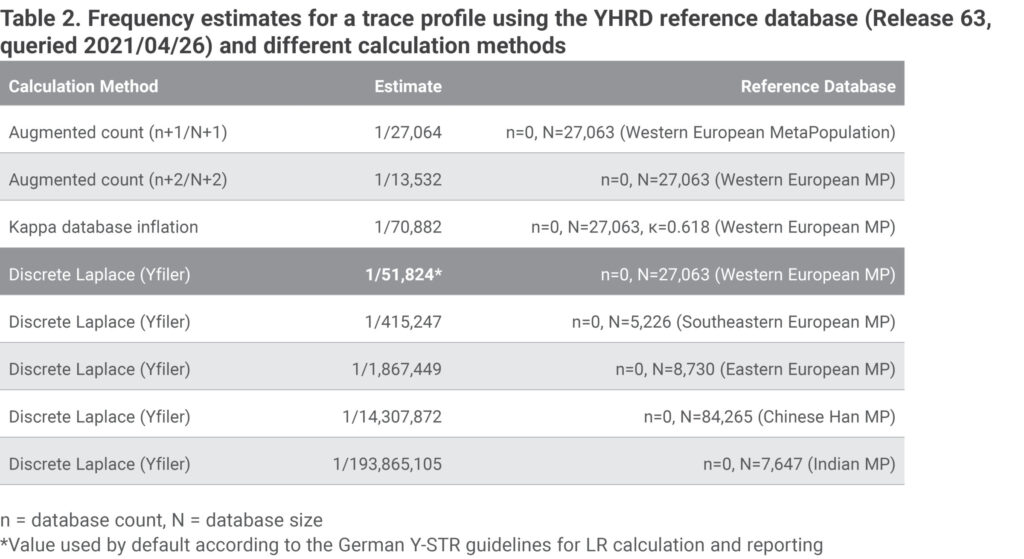
In Table 2 we compare different statistical values (reanalyzed using current YHRD release 63) for a trace/person match in a homicide case from Germany, which is described in detail in [10]. In accordance with the German Y-STR guidelines the DL value for the Western European metapopulation was used by default for reporting the results. The suspect was sentenced to life imprisonment and the ruling was confirmed by the highest appellate court in Germany.
The new guidelines mentioned earlier will help forensic practitioners in their daily life. But the evaluation of Y-STRs remains challenging and often the results are not very meaningful. However, with the understanding that ancestry is key for the interpretation of a Y match, it is possible to restructure databases and apply methods based on an evolutionary genetic model. So equipped, we expect growing acceptance of the Y-STR method within the forensic community, law enforcement and justice.
References
- Willuweit S, Anslinger K, Bäßler G, Eckert M, Fimmers R, Hohoff C, Kraft M, Leuker C, Molsberger G, Pich U, Razbin S, Rothämel T, Schneider H, Schneider PM, Templin M, Vennemann M, Wächter A, Weirich V, Zierdt H, Roewer L (2018) Joint recommendations of the project group “Statistical analysis of DNA” and the German Stain Commission on the statistical analysis of Y‑chromosomal DNA typing results. Rechtsmedizin 28(2)138-42.
- Rębała K, Branicki W, Pawłowski R, Spólnicka M, Kupiec T, Parys-Proszek A, Woźniak M, Grzybowski T, Boroń M, Wróbel M, Ciesielka M, Ossowski A, Jacewicz R (2020) Recommendations of the Polish Speaking Working Group of the International Society for Forensic Genetics on forensic Y chromosome typing. Archives of Forensic Medicine and Criminology 70 (1): 1–18.
- Rodriguez JJRB, Laude RP, De Ungria MCA (2021) An integrated system for forensic DNA testing of sexual assault cases in the Philippines. Forensic Sci Int Synerg 11 (3) 100133
- Forensic Science Regulator, Y- STR Profiling. FSR-G-227. Issue 1 [Online]. Available at: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/973580/227_Y_STR_guidance__Issue1.0_Final.pdf
- Roewer L, Andersen MM, Ballantyne J, Butler JM, Caliebe A, Corach D, D’Amato ME, Gusmão L, Hou Y, de Knijff P, Parson W, Prinz M, Schneider PM, Taylor D, Vennemann M, Willuweit S (2020) DNA Commission of the International Society of Forensic Genetics (ISFG): Recommendations on the Interpretation of Y-STR results in Forensic Analysis. Forensic Sci Int Genet. 48:102308.
- Andersen MM, Eriksen PS, Morling N (2013) The discrete Laplace exponential family and estimation of Y-STR haplotype frequencies. J Theor Biol. 329:39-51.
- Brenner CH (2010) Fundamental problem of forensic mathematics-the evidential value of a rare haplotype. Forensic Sci Int Genet 4(5):281-91.
- Balding D (2005) Weight-of-evidence for Forensic DNA. John Wiley & Sons Ltd.
- Willuweit S, Roewer L (2015) The new Y Chromosome Haplotype Reference Database. Forensic Sci Int Genet 15:43-8.
- Roewer L (2019) Y‐chromosome short tandem repeats in forensics – Sexing, profiling, and matching male DNA. Wiley Interdisciplinary Reviews 1(4) e1336
WOULD YOU LIKE TO SEE MORE ARTICLES LIKE THIS? SUBSCRIBE TO THE ISHI BLOG BELOW!
SUBSCRIBE NOW!