Variability in the amount of DNA deposited by individuals when touching a surface poses a significant hurdle to the evaluation of forensic analysis methods for touch samples. No two human latent fingerprint samples are the same due to the “shedder” status of different individuals as well as other factors including hand washing or deposition conditions.
Renee Williams, Signature Science
Generally, this limitation forces laboratories to utilize qualitative metrics (e.g., the frequency of successful analysis defined by the generation of a reportable DNA profile) rather than quantitative metrics (e.g., evaluating the total DNA recovery per sample) when assessing sample preparation methods or establishing proficiency among analysts.
To address these limitations, Signature Science has developed DNA-Touch™. This proprietary method uses synthetic fingermarks to accurately and consistently represent the biochemical composition and biological signatures of human fingerprints.
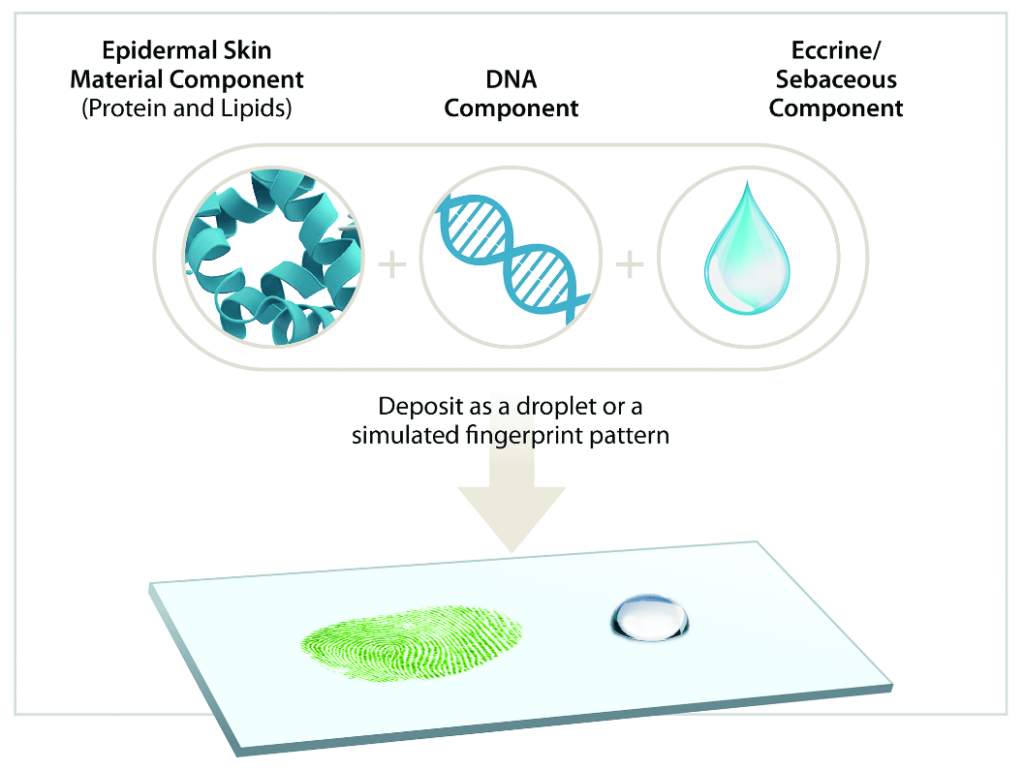
DNA-Touch samples include the primary components of a typical latent fingerprint, specifically sebaceous fluid, eccrine perspiration, extracellular DNA, and proteinaceous epidermal skin material (i.e., shed skin cells). An emulsion of synthetic sebaceous and eccrine material provides a chemically-relevant carrier solution into which a precise amount of extracted human genomic DNA is added.
This accurately mimics the extracellular DNA content of a typical latent print. Further, these samples also contain a representative quantity of protein originating from epidermal skin cells. This allows DNA-Touch to be used in combination with protein-based forensic identification techniques.
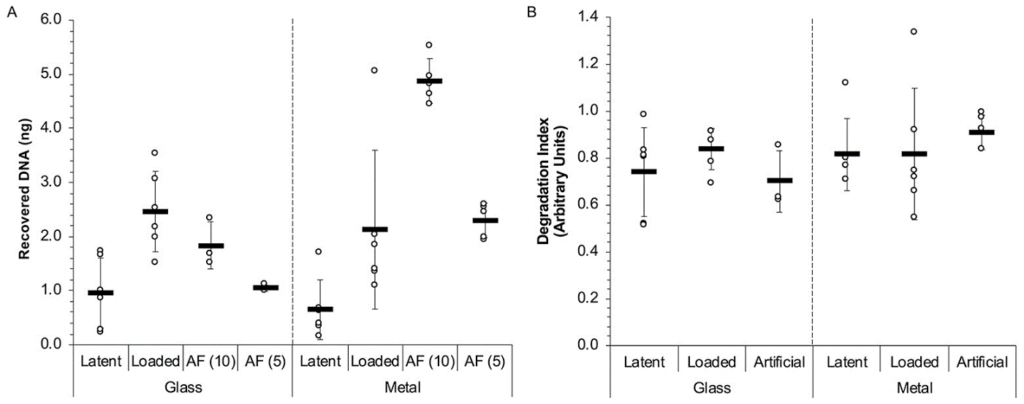
DNA-Touch samples can support the quantitative evaluation of laboratory performance (e.g. DNA or protein recovery, quantitation and typing success). Published data shows that DNA from the DNA-Touch samples (previously termed “artificial fingerprints” or “AFs”) was successfully recovered from glass and metal surfaces in ranges that generally mimicked the DNA yield ranges collected from human fingerprint samples. Evaluation of the degradation index revealed that the DNA was of comparable quality as well. Critically, the variability in extracted DNA yields between DNA-Touch replicates was lower than that of the human deposits. This illustrates the benefit of using touch samples containing a known, repeatable amount of genetic material when evaluating method or analyst performance.
“We view DNA-Touch as a game-changing technology for the analysis of touch forensic samples,” said Dr. Curt Hewitt, Signature Science’s Center for Advanced Genomics’ Science and Technology Advisor and study co-author. “While synthetic touch samples will never entirely replace human samples in forensic labs, DNA-Touch will both simplify and add scientific value to method development and validation efforts. The days of collecting dozens of latent fingerprints from multiple donors in the hopes of generating statistically defensible results for new collection methods are at an end.”
Envisioned applications for DNA-Touch samples in the forensic laboratory include staff training, measuring the competency and proficiency of both manual and automated extraction processes, and assessing value for emerging R&D and technology advancements. Signature Science’s Center for Advanced Genomics R&D team is partnering with forensic and R&D laboratories to refine and advance the technology. Please contact us if you are interested in evaluating this technology or joining our ‘early access’ community.
WOULD YOU LIKE TO SEE MORE ARTICLES LIKE THIS? SUBSCRIBE TO THE ISHI BLOG BELOW!
SUBSCRIBE NOW!